| AI in Clinical Medicine, ISSN 2819-7437 online, Open Access |
| Article copyright, the authors; Journal compilation copyright, AI Clin Med and Elmer Press Inc |
| Journal website https://aicm.elmerpub.com |
Review
Volume 2, April 2026, e21
Integrating Artificial Intelligence Into Sepsis Care: A Narrative Review of Predictive Models and Implementation Pathways
Abdul Alia , Hafiza Nabia
, Sher M. Sethib, e
, Arikah Minhajc
, Hadi Aziz Ansarid
, Ainan Arshada
aAga Khan University Hospital, Karachi, Pakistan
bDepartment of Medicine, Aga Khan University Hospital, Karachi, Pakistan
cJinnah Sindh Medical University, Karachi, Pakistan
dMedical College, Aga Khan University Hospital, Karachi, Pakistan
eCorresponding Author: Sher M. Sethi, Department of Medicine, Aga Khan University Hospital, Karachi 74800, Pakistan
Manuscript submitted March 17, 2026, accepted April 1, 2026, published online April 9, 2026
Short title: Sepsis Prediction With AI
doi: https://doi.org/10.14740/aicm21
| Abstract | ▴Top |
Background: Sepsis is a leading cause of morbidity and mortality, particularly in intensive care units (ICUs). Traditional prediction tools such as systemic inflammatory response syndrome (SIRS), Sequential Organ Failure Assessment (SOFA), and Acute Physiology and Chronic Health Evaluation II (APACHE II) are widely used but have limited specificity, responsiveness, and adaptability. This review evaluates artificial intelligence (AI)-based sepsis prediction models compared with conventional approaches, focusing on predictive performance, clinical integration, and future implementation pathways.
Methods: This narrative review was conducted using a structured literature search of PubMed, Scopus, and Web of Science databases for studies published between 2015 and 2025. Inclusion criteria comprised studies evaluating AI-based sepsis prediction models with reported performance metrics in adult or ICU settings. Approximately 40–50 studies were included after screening titles, abstracts, and references. Data were synthesized qualitatively with emphasis on model performance, validation, and clinical applicability.
Results: AI-based models generally outperformed traditional tools, with random forest, InSight, and DeepAISE achieving area under the receiver operating characteristic curve (AUROC) values above 0.90, superior sensitivity, and greater adaptability in dynamic ICU environments. Targeted Real-time Early Warning System (TREWS) demonstrated improved patient outcomes in prospective multi-site trials, enabling earlier antibiotic administration by several hours and associated reductions in mortality, highlighting meaningful real-world clinical benefit. Models such as Multi-task Gaussian Process Temporal Convolutional Network (MGP-TCN) further enhanced predictive accuracy using temporal data but remain computationally intensive. Integration into electronic health records and the potential for explainable AI were highlighted as key facilitators for clinical adoption.
Conclusions: AI-driven sepsis prediction models show significant advantages over conventional scoring systems, offering improved accuracy, adaptability, and workflow integration. However, widespread adoption requires multicenter prospective validation, enhanced interpretability to build clinician trust, and supportive policy frameworks to ensure safe, equitable, and sustainable implementation in clinical practice.
Keywords: Artificial intelligence; Health services; Intensive care; Predictive models; Sepsis
| Introduction | ▴Top |
Sepsis, defined as life-threatening organ dysfunction caused by a dysregulated host response to infection, remains a major global health challenge [1]. In 2017, it was estimated to account for 11 million deaths worldwide nearly 20% of all global deaths [2]. Despite advances in critical care, sepsis continues to be associated with high morbidity and mortality, with in-hospital death rates ranging from 10% to over 50% depending on severity and comorbidities [3].
To aid clinicians in early recognition and risk stratification, several scoring systems have been developed. The systemic inflammatory response syndrome (SIRS) criteria, while sensitive, lacks specificity and often leads to overdiagnosis. The Sequential Organ Failure Assessment (SOFA) and its simplified variant qSOFA provide better prognostic accuracy but require extensive laboratory data, limiting their utility for rapid assessment. Similarly, the Acute Physiology and Chronic Health Evaluation II (APACHE II) score is widely applied but remains complex and time-consuming in urgent settings [4, 5]. Collectively, these tools face challenges of low specificity, delayed responsiveness, and limited adaptability in dynamic intensive care unit (ICU) environments.
Emerging artificial intelligence (AI)-based algorithms offer a promising alternative by integrating large, multidimensional clinical datasets for timely and accurate sepsis prediction. Models such as random forest, InSight, and DeepAISE have demonstrated area under the receiver operating characteristic curve (AUROC) values > 0.9 in both internal and external validations, consistently outperforming conventional tools [6, 7]. More recently, prospective evaluation of the Targeted Real-time Early Warning System (TREWS) showed improved outcomes through earlier antibiotic initiation [8]. In addition, the first Food and Drug Administration (FDA)-authorized AI/machine learning (ML) tool for sepsis prediction was approved in 2024, reflecting the translational potential of these systems in clinical practice [9].
Given the persistent diagnostic challenges of sepsis and the growing body of evidence supporting AI-driven prediction models, this review synthesizes current developments, evaluates clinical applicability, and discusses future directions for integrating AI into sepsis management.
| Methods | ▴Top |
We conducted a narrative review of published literature on AI-driven sepsis prediction models. A structured search of PubMed, Scopus, and Web of Science was performed for studies published between January 2015 and December 2025. Search terms included combinations of “sepsis,” “artificial intelligence,” “machine learning,” “deep learning,” and “prediction models.”
Studies were included if they: 1) evaluated AI/ML-based models for sepsis prediction, 2) reported performance metrics such as AUROC, sensitivity, or specificity, and 3) involved adult or ICU populations. Studies focusing solely on pediatric populations, non-clinical simulations, or lacking performance metrics were excluded.
Approximately 40–50 relevant studies were reviewed, including retrospective, prospective, and validation studies. Key model characteristics, validation approaches, and performance metrics were extracted and qualitatively synthesized.
| Results | ▴Top |
AI models for sepsis prediction
In recent years, AI has been increasingly applied to sepsis prediction, leading to the development of diverse models aimed at early detection and timely intervention [10]. These models vary in methodology, input requirements, and clinical validation, yet most consistently outperform traditional clinical scoring systems. Below we summarize key AI models currently in use or under study.
Epic Sepsis Model (ESM)
The ESM, developed by Epic Systems, is among the most widely deployed proprietary sepsis prediction tools in the United States. Released in 2018, it has been implemented in hundreds of hospitals. However, external validation has revealed significant shortcomings. A large study by Wong et al reported that ESM achieved an AUROC of only 0.63, substantially lower than expected, with high false-positive rates and poor calibration. Its lack of transparency (a “black box” model) and reliance on institution-specific training data are believed to contribute to poor generalizability and clinician distrust [11].
InSight
Developed by University of California, San Francisco (UCSF) and the University of Chicago, InSight is a lightweight ML model requiring only six routinely collected vital signs. First published in 2016, it demonstrated robust cross-institutional validity with AUROC values consistently exceeding 0.90, outperforming SIRS, Modified Early Warning Score (MEWS), and qSOFA. Its minimal input requirements make it especially suitable for settings with sparse or incomplete data [6].
Sepsis Watch
Sepsis Watch, developed by Duke University Health System in collaboration with HBI Solutions, employs deep learning via recurrent neural networks (RNNs) to predict sepsis risk in emergency department (ED) patients. Launched in 2019, it integrates real-time electronic health record (EHR) data and was prospectively validated in ED settings. In one evaluation, it demonstrated AUROC values between 0.85 and 0.90, showing improved early recognition compared to clinician judgment alone [12]. Its major strength lies in workflow integration, although reliance on complex RNNs may limit interpretability.
DeepAISE
DeepAISE is a deep learning–based recurrent neural survival model developed by researchers at UC San Diego and Emory University in 2021. It processes 65 routinely collected variables to generate hourly risk scores beginning 4 h after ICU admission. In validation studies, DeepAISE achieved AUROC values of 0.90 (internal) and 0.87 (external), with the added advantage of interpretability by highlighting key contributing factors for individual patients [7].
TREWS
Developed at Johns Hopkins University, TREWS combines rule-based and ML methods. Notably, it is among the first AI models validated in a prospective, multi-site clinical trial, where it demonstrated earlier antibiotic initiation and improved outcomes [8]. This real-world validation highlights its translational potential, though it remains limited to health systems with robust EHR integration.
MGP-TCN
The Multi-task Gaussian Process Temporal Convolutional Network (MGP-TCN), introduced in 2019, leverages time-series ICU data to capture complex temporal patterns. Subsequent enhancements, such as the GRU-D-MGP-TCN hybrid, have further improved predictive performance, achieving AUROC values up to 0.99 and area under the precision–recall curve (AUPRC) of 0.96 [13]. While technically advanced, these models are computationally intensive and not yet widely implemented in clinical practice. Table 1 illustrates the comparison of different AI models.
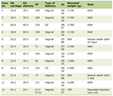 Click to view | Table 1. Comparative Summary of AI Models for Sepsis Prediction |
| Application of AI Model in Sepsis Prediction in ICU Setting | ▴Top |
Sepsis progression in critically ill patients is highly dynamic, and delayed recognition is strongly associated with increased mortality. Early identification reduces in-hospital mortality from over 20% to < 10% in some cohorts [14]. Traditional tools such as SIRS, SOFA, and qSOFA provide variable predictive performance depending on infection site and patient population, limiting their reliability in diverse ICU settings [15].
AI-based models have shown promise in addressing these limitations by integrating high-dimensional data, handling irregular sampling, and producing timely alerts. Their performance is typically measured using AUROC, a measure of overall discriminative ability and AUPRC, which is especially relevant in imbalanced datasets like sepsis. For context, AUROC values above 0.85 are considered strong, while AUPRC reflects precision in detecting true positives among many negatives.
DeepAISE
DeepAISE is a recurrent neural survival model trained on 65 routine clinical variables, generating hourly risk scores from ICU admission onward. In validation studies, it achieved AUROC 0.90 (internal) and 0.87 (external), with sensitivity 0.85 and specificity 0.80 [7]. Unlike many black-box systems, it identifies key contributing factors in real-time, improving interpretability and clinical trust.
MGP-TCN and hybrid models
The MGP-TCN handles irregular time-series ICU data and models temporal dynamics. Initial implementations improved AUROC from baseline values (∼0.75) to 0.91, with AUPRC 0.69 [14]. More recently, the hybrid GRU-D-MGP-TCN achieved AUROC 0.99 and AUPRC 0.96, representing near-perfect discrimination in retrospective ICU datasets [13].
InSight
InSight is a lightweight ML model using only six vital signs, making it robust in settings with missing or sparse data. Multicenter validations across UCSF, Stanford, BIDMC, and community hospitals demonstrated AUROC consistently above 0.90, outperforming SIRS, MEWS, and qSOFA [6]. Its simplicity and generalizability highlight its potential for wider ICU adoption, including resource-limited environments.
While these models demonstrate strong predictive accuracy, integration into ICU workflows remains challenging. Their performance depends on local data quality, EHR compatibility, and clinician acceptance. Furthermore, frequent false alerts may reduce trust and lead to underutilization. Hence, successful implementation requires not only technical accuracy but also seamless workflow integration, interpretability, and prospective validation [6, 7, 14]. Figure 1 shows the workflow of AI-sepsis prediction in ICU.
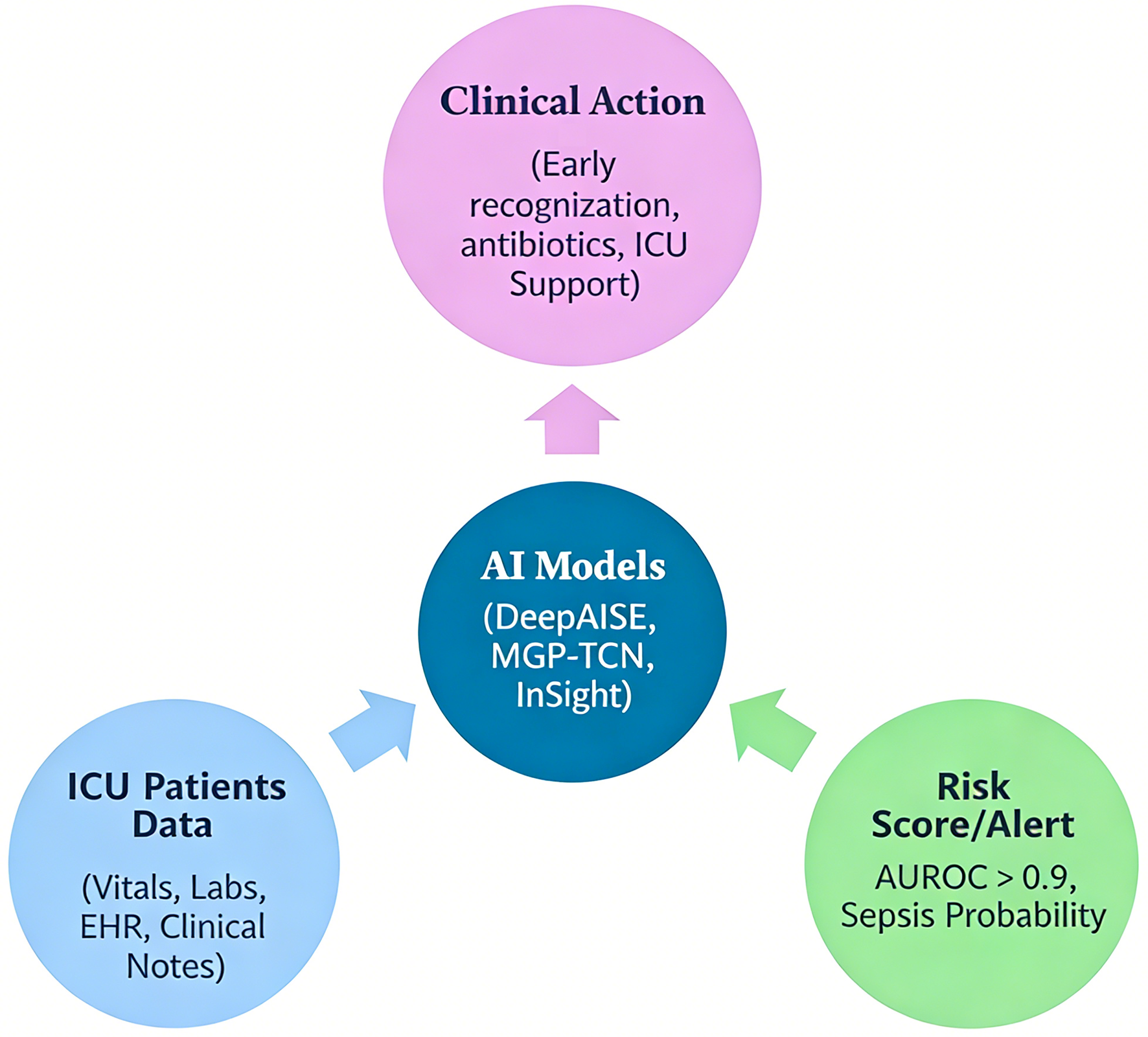 Click for large image | Figure 1. Workflow of artificial intelligence (AI)-based sepsis prediction in the intensive care unit (ICU). This figure illustrates the end-to-end pipeline of AI-driven sepsis prediction. Patient data, including vital signs, laboratory values, and electronic health record (EHR) inputs, are continuously collected and pre-processed (data cleaning, normalization, and handling of missing values). These data are then fed into machine learning or deep learning models (e.g., random forest, recurrent neural networks, or temporal convolutional networks), which generate dynamic risk scores in real time. The output is integrated into clinical workflows via EHR-based alert systems, prompting early clinical evaluation and intervention (e.g., antibiotic initiation, fluid resuscitation). The figure also highlights feedback loops for model refinement and the role of clinician oversight to ensure safe and effective implementation. |
| Comparison of AI Model to Conventional Sepsis Prediction Tools (SIRS, SOFA, APACHE) | ▴Top |
Modern AI/ML models (random forest, DeepAISE, InSight, and temporal networks such as MGP-TCN) demonstrate consistently higher discrimination for early sepsis prediction than conventional tools (SIRS, qSOFA, SOFA, APACHE II). For example, DeepAISE reports AUROC: 0.90, InSight achieves AUROC: 0.88–0.92 across multicenter evaluations, and temporal models (MGP-TCN and hybrids) report AUROC: 0.90 or higher depending on cohort and horizon. In contrast, commonly used diagnostic criteria such as SIRS, qSOFA, and APACHE show lower specificity and lower AUROC in many cohorts [5, 15, 16]. Table 2 shows the comparison of AI models with conventional sepsis prediction tools.
 Click to view | Table 2. Comparison of AI Models With Conventional Sepsis Prediction Tools |
Validation of AI Sepsis Prediction Model
Validation is essential to determine the accuracy, generalizability, and clinical applicability of AI models. Three major stages are typically recognized: internal validation (performance within the training dataset), external validation (testing in independent datasets or institutions), and prospective validation (evaluation in real-world clinical practice). Most published AI models for sepsis prediction have undergone retrospective internal and external validation, demonstrating AUROC values > 0.85 in multiple cohorts. However, prospective trials remain rare but critical, as they provide evidence of real-world impact on patient outcomes and clinical workflows. For example, TREWS is one of the few AI systems tested in a multi-site prospective trial, where it enabled earlier antibiotic administration and reduced mortality [8].
From a regulatory perspective, rigorous validation and transparent reporting are prerequisites for clinical adoption. The US FDA and European Medicines Agency (EMA) require robust evidence of safety, efficacy, and generalizability before authorizing AI/ML-based clinical tools. In 2020, the CONSORT-AI extension was introduced to improve transparency and reporting standards in trials involving AI interventions, ensuring that studies describe datasets, model versions, validation approaches, and interpretability features [17]. More recently, the FDA authorized its first AI/ML tool for sepsis prediction [18], highlighting a growing emphasis on regulatory oversight.
Thus, while retrospective validation provides proof of concept, prospective, multicenter clinical trials and adherence to regulatory standards are essential steps to establish trust, ensure reproducibility, and enable the safe integration of AI into sepsis care. Figure 2 demonstrates the validation hierarchy of AI sepsis model.
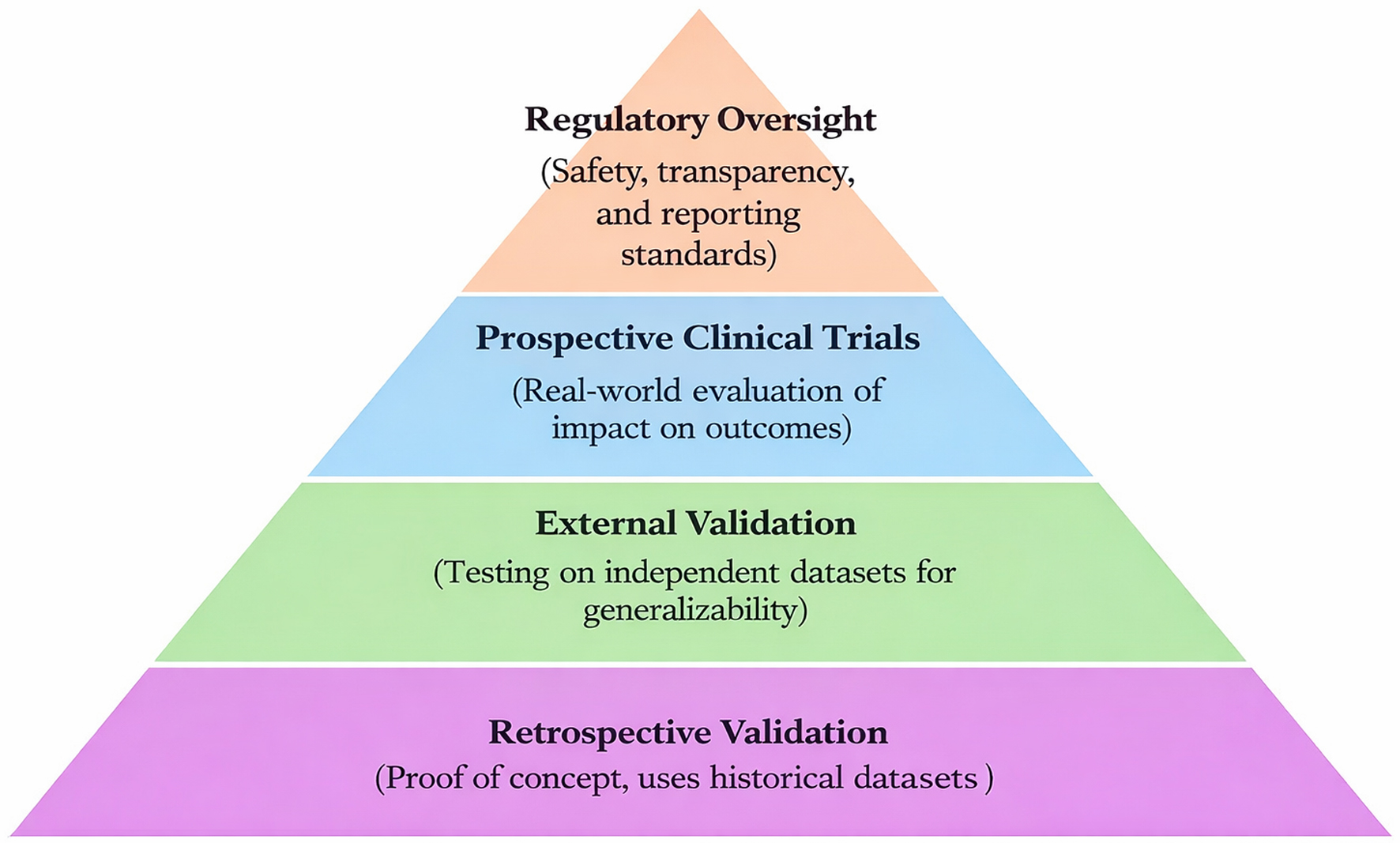 Click for large image | Figure 2. Validation hierarchy of artificial intelligence (AI)-based sepsis prediction models. This figure depicts the sequential stages of validation required for AI models prior to clinical implementation. Internal validation evaluates model performance within the training dataset using techniques such as cross-validation. External validation assesses generalizability across independent datasets or institutions with different patient populations and clinical settings. Prospective validation represents real-world testing in clinical environments, measuring impact on patient outcomes, workflow integration, and decision-making. The figure emphasizes that while many models demonstrate strong performance in retrospective analyses, relatively few have undergone prospective, multicenter validation. Regulatory oversight and reporting standards (e.g., CONSORT-AI) are also highlighted as essential components for safe clinical adoption. |
| Challenges and Limitations | ▴Top |
Despite promising results, several challenges limit the widespread adoption of AI in sepsis prediction.
Data quality and heterogeneity
Real-world EHR data are often incomplete, irregular, and inconsistently documented, which can degrade model performance outside controlled research settings. In addition, variability in sepsis definitions (sepsis-2 vs. sepsis-3) introduces labeling inconsistencies that complicate training and validation.
Generalizability
Models trained on institution-specific data frequently show performance declines when applied in new healthcare systems due to differences in patient demographics, disease patterns, workflows, and technical infrastructure. This “dataset shift” remains a critical barrier to reliable deployment.
Clinical integration and trust
Many AI models are perceived as “black boxes,” providing risk scores without clear explanations. This lack of interpretability reduces clinician confidence and may lead to underuse, especially when alerts are triggered before overt clinical deterioration. Workflow disruption and false alarms can further contribute to alert fatigue.
Ethical and equity concerns
AI models may inadvertently perpetuate or amplify healthcare disparities if trained on biased datasets. Ensuring fairness, transparency, and equitable performance across diverse patient populations is essential to avoid widening gaps in care.
| Future Direction and Innovation | ▴Top |
As AI continues to evolve, future innovations in sepsis prediction should focus on clinical relevance, generalizability, and trust. Real-time integration into EHR systems remains a priority, enabling seamless incorporation into existing workflows. Personalized prediction models that adapt to patient-specific baselines and continuous learning frameworks could further improve accuracy and clinical utility.
Federated learning offers a promising approach to enhance generalizability by training models on decentralized datasets without compromising patient privacy. At the same time, explainable AI is essential to build clinician trust by providing transparent reasoning behind predictions, supporting human–AI collaboration rather than replacement.
The fusion of multimodal data sources including vital signs, clinical notes, imaging, wearables, and genomics may uncover novel sepsis phenotypes and enable even earlier interventions. However, prospective multicenter trials remain critical to establish true clinical benefit, as demonstrated by TREWS. Regulatory frameworks, such as those promoted by the FDA and CONSORT-AI guidelines, will be essential to ensure safety and accountability.
Finally, collaboration between clinicians, data scientists, health system leaders, and policymakers will be crucial to translate technical advances into real-world practice. Aligning innovation with ethical and operational frameworks will help ensure equitable, scalable, and sustainable deployment of AI tools for sepsis care.
| Discussion | ▴Top |
The integration of AI into sepsis prediction represents a paradigm shift in critical care medicine, with the potential to overcome many of the limitations of traditional scoring systems such as SIRS, SOFA, and APACHE II [4, 5]. In this review, AI-driven models including InSight, DeepAISE, TREWS, and MGP-TCN demonstrated superior predictive accuracy, achieving AUROC values above 0.90 in several studies [6–8, 13]. These findings highlight the enhanced capability of ML algorithms to process complex, high-dimensional datasets and capture temporal relationships that are often missed by rule-based systems. Such adaptability is particularly valuable in the dynamic intensive care environment, where clinical deterioration can occur rapidly and unpredictably.
The prospective validation of the TREWS marked a crucial milestone, demonstrating improved patient outcomes through earlier antibiotic administration (often several hours before clinical recognition) and associated reductions in mortality, supporting real-world clinical impact [8]. This supports the premise that predictive precision can translate into tangible clinical benefit when seamlessly embedded within routine workflows. Nonetheless, the generalizability of AI models remains a major challenge. Many systems are developed using single-center or proprietary datasets, leading to model drift and performance degradation when applied to different patient populations or EHR infrastructures [11, 13]. To ensure reproducibility and real-world applicability, future research must emphasize multicenter, prospective trials with standardized sepsis definitions and transparent model reporting, as recommended by the CONSORT-AI guidelines [17].
Interpretability and clinician trust are equally vital for successful adoption. Although black-box models such as deep learning networks demonstrate remarkable accuracy, their lack of transparency can hinder clinical confidence and reduce the likelihood of adoption [7, 11]. Explainable AI approaches, exemplified by DeepAISE, offer promising strategies by identifying the relative contribution of clinical features to each prediction, thus aligning algorithmic reasoning with human decision-making [7]. Such interpretability not only enhances usability but also supports ethical and regulatory compliance by clarifying accountability in AI-assisted care.
The recent authorization of an AI/ML-based sepsis prediction tool by the US FDA underscores growing recognition of AI’s clinical potential while emphasizing the necessity of robust validation and continuous post-deployment monitoring [9]. Regulatory frameworks are increasingly focusing on transparency, algorithmic fairness, and bias mitigation to ensure equitable outcomes across diverse populations. Ethical considerations particularly around data privacy, representativeness, and explainability must remain central to AI system development and deployment to prevent the amplification of existing healthcare disparities [10, 17].
However, key limitations include limited prospective and external validation across diverse healthcare settings, dataset heterogeneity affecting generalizability, and the high computational requirements of advanced deep learning models, which may restrict real-world implementation.
Moving forward, innovations such as federated learning and multimodal data integration (combining vital signs, laboratory data, clinical notes, imaging, and genomics) are expected to enhance generalizability and personalization of sepsis prediction models [17, 18]. However, these advances will require robust computational infrastructure and cross-disciplinary collaboration among clinicians, data scientists, informaticians, and policymakers. Ultimately, the transformative potential of AI in sepsis management will depend not only on algorithmic excellence but also on human-centered design, transparent governance, and sustainable clinical integration strategies.
Acknowledgments
None to declare.
Financial Disclosure
There was no funding or institutional support for this study.
Conflict of Interest
The authors declare there is no conflict of interest to disclose.
Author Contributions
AA: conceptualization; designing; drafting the article. HN: literature search and data acquisition; drafting the article. SS: conceptualization; designing; reviewing the article. AM: literature search and data acquisition; drafting the article. HA: literature search and data acquisition; drafting the article. AAr: reviewing the article, proof-reading, supervision.
Data Availability
All data generated for this study are included in this review article. Further information if required can be obtained from corresponding author.
| References | ▴Top |
- Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M, Bellomo R, et al. The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA. 2016;315(8):801-810.
doi pubmed - Saito H. Sepsis in Global Health: Current global strategies to fight against sepsis. Acute Med Surg. 2025;12(1):e70045.
doi pubmed - Long B, Gottlieb M. Emergency medicine updates: Management of sepsis and septic shock. Am J Emerg Med. 2025;90:179-191.
doi pubmed - Li Y, Yan C, Gan Z, Xi X, Tan Z, Li J, Li G. Prognostic values of SOFA score, qSOFA score, and LODS score for patients with sepsis. Ann Palliat Med. 2020;9(3):1037-1044.
doi pubmed - Koch C, Edinger F, Fischer T, Brenck F, Hecker A, Katzer C, Markmann M, et al. Comparison of qSOFA score, SOFA score, and SIRS criteria for the prediction of infection and mortality among surgical intermediate and intensive care patients. World J Emerg Surg. 2020;15(1):63.
doi pubmed - Desautels T, Calvert J, Hoffman J, Jay M, Kerem Y, Shieh L, Shimabukuro D, et al. Prediction of Sepsis in the Intensive Care Unit With Minimal Electronic Health Record Data: A Machine Learning Approach. JMIR Med Inform. 2016;4(3):e28.
doi pubmed - Shashikumar SP, Josef CS, Sharma A, Nemati S. DeepAISE - An interpretable and recurrent neural survival model for early prediction of sepsis. Artif Intell Med. 2021;113:102036.
doi pubmed - Adams R, Henry KE, Sridharan A, Soleimani H, Zhan A, Rawat N, Johnson L, et al. Prospective, multi-site study of patient outcomes after implementation of the TREWS machine learning-based early warning system for sepsis. Nat Med. 2022;28(7):1455-1460.
doi pubmed - Bhargava A, Lopez-Espina C, Schmalz L, et al. FDA-Authorized AI/ML tool for sepsis prediction: development and validation. NEJM AI. 2024;1(12):AIoa2400867.
doi - Wu M, Du X, Gu R, Wei J. Artificial Intelligence for Clinical Decision Support in Sepsis. Front Med (Lausanne). 2021;8:665464.
doi pubmed - Wong A, Otles E, Donnelly JP, Krumm A, McCullough J, DeTroyer-Cooley O, Pestrue J, et al. External Validation of a Widely Implemented Proprietary Sepsis Prediction Model in Hospitalized Patients. JAMA Intern Med. 2021;181(8):1065-1070.
doi pubmed - Sendak MP, D’Arcy J, Kashyap S, et al. A path for translation of machine learning products into healthcare delivery. EMJ Innov. 2020;10:19-00172.
doi - Moor M, Horn M, Rieck B, et al. Early recognition of sepsis with Gaussian process temporal convolutional networks and dynamic time warping. InMachine learning for healthcare conference. PMLR. 2019; p. 2-26.
doi - Jee W, Jo S, Lee JB, Jin Y, Jeong T, Yoon JC, Park B. Mortality difference between early-identified sepsis and late-identified sepsis. Clin Exp Emerg Med. 2020;7(3):150-160.
doi pubmed - Zhang D, Wang C, Li Q, Zhu Y, Zou H, Li G, Zhan L. Predictive value of multiple scoring systems in the prognosis of septic patients with different infection sites: analysis of the medical information mart for the intensive care IV database. Biomedicines. 2024;12(7):1415.
doi pubmed - Gao J, Lu Y, Ashrafi N, Domingo I, Alaei K, Pishgar M. Prediction of sepsis mortality in ICU patients using machine learning methods. BMC Med Inform Decis Mak. 2024;24(1):228.
doi pubmed - Bignami EG, Berdini M, Panizzi M, Domenichetti T, Bezzi F, Allai S, Damiano T, et al. Artificial Intelligence in Sepsis Management: An Overview for Clinicians. J Clin Med. 2025;14(1).
doi pubmed - Lee S, Shin G, Hwang J, et al. Early prediction of sepsis in the intensive care unit using the GRU-D-MGP-TCN model. IEEE Access. 2024.
doi
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, including commercial use, provided the original work is properly cited.
AI in Clinical Medicine is published by Elmer Press Inc.